TL;DR:
- Proper early selection and validation of molecular assays are crucial for regulatory compliance and data integrity.
- Different assay platforms like PCR, NGS, and immunoassays serve distinct purposes with unique advantages and limitations.
- Advanced techniques and AI integration are expanding capabilities but require rigorous validation and lifecycle management.
Selecting the wrong molecular assay can stall a drug development program by months, trigger costly regulatory resubmissions, or invalidate years of preclinical data. Yet many pharmaceutical and biomedical teams still treat assay selection as a downstream decision rather than a foundational one. The reality is that your assay strategy shapes everything from data quality to regulatory package strength. This guide covers the core assay types, validation frameworks, and emerging technologies your team needs to make confident, compliant decisions at every stage of product development.
Table of Contents
- What is a molecular assay and why does it matter?
- Common molecular assay methods: An in-depth overview
- Validation and regulatory demands for molecular assays
- Advanced molecular assay strategies and emerging challenges
- What most guides miss about molecular assays
- How Materials Metric accelerates your assay programs
- Frequently asked questions
Key Takeaways
| Point | Details |
|---|---|
| Foundational definitions | Molecular assays are essential analytical tools for detecting and quantifying biomolecules in pharmaceutical and biomedical R&D. |
| Choose methods wisely | Selecting the right assay method depends on your research goal, regulatory endpoint, and sample type. |
| Validation is crucial | Comprehensive validation ensures regulatory acceptance and robust data for clinical or exploratory use. |
| Embrace advanced strategies | Cutting-edge approaches like cfDNA NGS and AI optimization expand assay possibilities but require careful risk management. |
| Orthogonal confirmation | Using multiple, complementary assays often yields stronger and more reliable results for product development and compliance. |
What is a molecular assay and why does it matter?
A molecular assay is an analytical technique used to detect, quantify, or characterize biomolecules such as nucleic acids, proteins, and biomarkers within a biological sample. As biomedical assay basics confirm, molecular assays detect and quantify biomolecules using methods like PCR, qPCR, NGS, HPLC, immunoassays, and cell-based assays. These platforms are not interchangeable. Each has distinct sensitivity thresholds, throughput profiles, and regulatory acceptance histories.
For pharmaceutical and biomedical companies, the stakes are high. Assays underpin every critical decision in your pipeline, from early target identification to post-market surveillance. A poorly selected or inadequately validated assay introduces variability that regulators will flag, and that your competitors will not have. Getting it right the first time is both a scientific and a business priority.

Our molecular assay development work consistently shows that teams who invest early in method selection avoid the most expensive rework cycles. The choice of platform must align with your product type, sample matrix, and the regulatory pathway you are pursuing.
One figure that captures the precision now achievable: NGS assays deliver sensitivity above 96% for single nucleotide variants (SNVs) and specificity exceeding 99.9999%, making them a powerful option for mutation profiling and biomarker discovery. Understanding where each method excels is the first step toward building a defensible assay strategy.
Top application areas for molecular assays:
- Pharmaceutical R&D and drug candidate profiling
- Companion diagnostic and biomarker discovery programs
- Biosimilar characterization and comparability studies
- Advanced therapy medicinal products (ATMPs) including gene and cell therapies
- Regulatory submission packages for IND, BLA, and NDA filings
For teams navigating assay methods and compliance, understanding these foundational distinctions is the prerequisite for everything that follows.
Common molecular assay methods: An in-depth overview
Every major assay platform has a specific niche. Choosing based on habit or lab availability rather than fit-for-purpose criteria is one of the most common and costly mistakes in pharmaceutical development. Key methodologies include PCR for nucleic acid amplification, chromatographic assays such as HPLC and GC for separation and quantification, spectroscopic methods like UV-Vis and FTIR, mass spectrometry, and biological and immunoassays.
| Method | Principle | Pros | Cons | Common use case |
|---|---|---|---|---|
| PCR/qPCR | Nucleic acid amplification | Fast, sensitive, low cost | Limited multiplexing | Pathogen detection, gene expression |
| NGS | Massively parallel sequencing | Broad profiling, high throughput | Complex data analysis | Mutation profiling, biomarker discovery |
| HPLC/Chromatography | Separation by physicochemical properties | Quantitative, reproducible | Requires pure samples | Small molecule drug quantification |
| Immunoassay (ELISA) | Antigen-antibody binding | High specificity, scalable | Matrix effects common | Protein quantification, pharmacokinetics |
| Cell-based assay | Functional cellular response | Reflects biological activity | Variable, labor intensive | Potency testing, mechanism of action |
Standard steps in a PCR-based molecular assay:
- Sample collection and nucleic acid extraction
- Primer and probe design for target specificity
- Thermal cycling for amplification
- Signal detection and quantification
- Data normalization against reference controls
- Statistical analysis and result interpretation
For broader genetic profiling, NGS remains the platform of choice. For high-accuracy single-target confirmation, Sanger sequencing still holds value. ELISA continues to be the workhorse for protein quantification in pharmacokinetic studies. Our molecular diagnostics lab supports all of these platforms with method development tailored to your product and regulatory context.
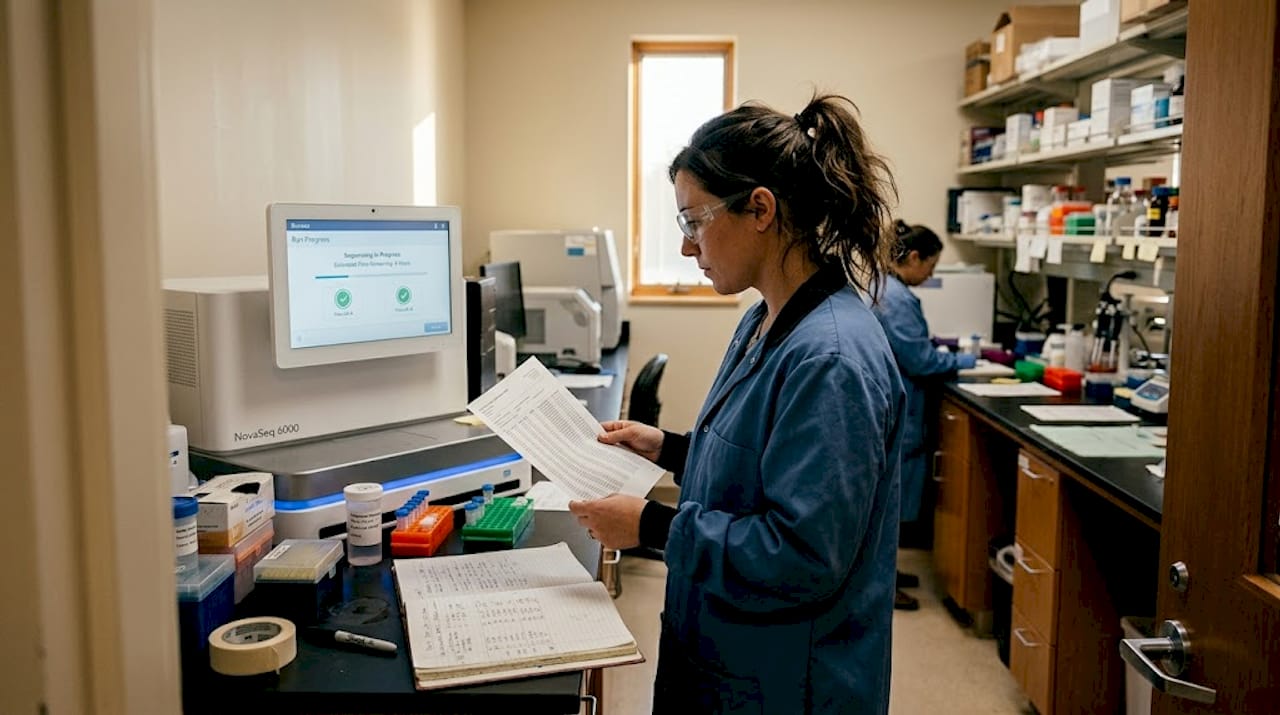

When evaluating screening assay approaches, the choice of detection chemistry matters as much as the instrument platform. Ligand-binding assays, for example, require careful attention to endogenous analyte interference. NMR in molecular assays adds a structural dimension that chromatographic and immunological methods cannot provide.
Pro Tip: Always confirm endogenous analyte recognition in ligand-binding assays before advancing to formal validation. Missed interference at this stage is a common cause of regulatory queries during clinical submissions.
Validation and regulatory demands for molecular assays
Validation is where scientific rigor meets regulatory expectation. Regulatory agencies do not accept assay data at face value. They require documented evidence that your method performs as claimed, under the conditions in which it will be used. The fit-for-purpose validation framework, codified in CLSI MM17, defines how validation depth should scale with intended use.
For risk-based validation guidelines, the principle is clear: full validation is required for assays supporting clinical decisions or regulatory submissions, while partial validation is acceptable for exploratory or preclinical work. Applying full validation resources to every early-stage assay wastes time and budget. Applying partial validation to a pivotal assay is a regulatory risk you cannot afford.
“Fit-for-purpose validation means the extent of validation is matched to the intended use of the assay, ensuring that resources are directed where the regulatory and clinical stakes are highest.” — CLSI MM17
Key validation parameters your team must document:
- Specificity: ability to distinguish target from non-target analytes
- Sensitivity and limit of detection (LOD): lowest reliably detected concentration
- Accuracy: agreement between measured and true values
- Precision: repeatability within and across runs
- Linearity and range: performance across expected sample concentrations
- Robustness: stability under minor method variations
Orthogonal validation is a strategy that many teams underuse. Running a complementary method alongside your primary assay, such as qPCR to confirm NGS variant calls, gives you cross-platform confidence and strengthens your regulatory package. Our custom assay validation programs are built around this principle.
For teams working through testing assay validation for the first time, the most important early investment is a validation plan that anticipates edge cases. Low variant allele fractions, complex sample matrices, and cross-reactive analytes should be stress-tested before you reach the regulatory submission stage.
Pro Tip: Address low variant allele fraction (VAF) scenarios and matrix effects during assay design, not after validation failures. Tailored in silico dilution experiments at the development stage are far less costly than protocol amendments during regulatory review.
Advanced molecular assay strategies and emerging challenges
The frontier of molecular assay development is moving fast. Combined RNA and DNA exome profiling, circulating tumor DNA (ctDNA) detection, and AI-assisted assay optimization are reshaping what is technically achievable. Advanced assays like combined RNA/DNA exome or cfDNA NGS enable comprehensive profiling of SNVs, fusions, tumor mutational burden (TMB), and microsatellite instability (MSI), supporting precision oncology research with positive percent agreement (PPA) and negative percent agreement (NPA) exceeding 95%.
| Assay type | Sensitivity | Specificity | LOD | Key challenge |
|---|---|---|---|---|
| cfDNA NGS (plasma) | >95% PPA | >99% NPA | 0.1% VAF | Low ctDNA fraction, fragmentation noise |
| RNA/DNA exome | >96% | >99.9% | SNV level | Data complexity, fusion detection |
| Targeted amplicon NGS | >98% | >99% | 1% VAF | PCR bias, primer design |
| Digital PCR (ddPCR) | >99% | >99.9% | 0.01% VAF | Throughput limitations |
For NGS and cfDNA insights, the key technical challenge is low ctDNA fraction in plasma samples. Low VAF detection in cfDNA is complicated by noise from fragmentation, low ctDNA fraction, and sample degradation, requiring deep sequencing, patient-matched references, and machine learning model tuning.
AI in assay optimization is no longer experimental. Machine learning models now assist in variant caller tuning, multiplexing panel design, and quality control flagging, reducing manual review burden and improving reproducibility across sites.
Current challenges in advanced molecular assay development:
- Multiplexing panel design without cross-reactivity
- Managing sample matrix effects in complex biological fluids
- Automating high-throughput workflows without sacrificing accuracy
- Tuning ML models for rare variant detection
- Maintaining cross-batch consistency in longitudinal studies
Cell-based assays remain essential alongside molecular platforms, particularly for ATMPs and biologics where functional activity must be demonstrated. Flow cytometry and reporter gene assays provide mechanistic evidence that sequence-level data alone cannot deliver. Combining molecular and precision oncology NGS approaches with functional readouts gives your regulatory package both breadth and depth.
What most guides miss about molecular assays
Most articles on molecular assays focus on the technology. They compare platforms, list sensitivity numbers, and move on. What they rarely address is the organizational and strategic gap that causes the most real-world failures: the neglect of validation lifecycle management and orthogonal confirmation.
We see this pattern regularly. A team adopts a next-generation NGS platform or an AI-assisted variant caller, invests heavily in the technology, and then underinvests in the validation infrastructure that makes the data defensible. The flashiest tool in the lab is worth nothing if the validation package cannot withstand regulatory scrutiny.
The uncomfortable truth is that cross-batch variation, reference sample handling, and reagent lot qualification are where most assay programs quietly fail. These are not glamorous problems. They do not feature in conference presentations. But they are the day-to-day bottlenecks that determine whether your data holds up at submission.
Our perspective is that validation-centric approaches should be designed in parallel with platform selection, not after it. Fit-for-purpose is not a shortcut. It is a disciplined framework that requires you to define intended use before you write a single validation protocol. Teams that do this well move faster, not slower, through regulatory review.
How Materials Metric accelerates your assay programs
Building a molecular assay program that satisfies both scientific rigor and regulatory expectations requires more than good instrumentation. It requires a partner who understands the full development lifecycle.
At Materials Metric, we work as an extension of your research and regulatory teams, supporting assay platform selection, method development, and validation strategy from early-stage discovery through submission-ready packages. Our capabilities span analytical testing impact across spectroscopy, chromatography, and molecular diagnostics, backed by ISO 9001:2015 and GLP-aligned quality systems. From material characterization techniques to integrated chemical and microscopy services, we bring the depth your program needs. Contact us to discuss your assay challenges and timelines.
Frequently asked questions
What is the key difference between qPCR and NGS assays?
qPCR quantifies specific nucleic acids quickly and cost-effectively for targeted applications, while NGS delivers higher throughput than PCR and broader genetic profiling across thousands of targets simultaneously.
How do regulatory agencies assess molecular assay validation?
Agencies require fit-for-purpose validation with documented evidence on specificity, sensitivity, accuracy, precision, and robustness, with validation depth scaled to the clinical or regulatory significance of the assay.
What challenges occur with cfDNA-based molecular assays?
Low VAF, fragmentation noise, and sample degradation reduce sensitivity and specificity in plasma-based cfDNA assays, requiring deep sequencing strategies and patient-matched reference controls to mitigate.
Why use both cell-based and molecular assays?
Combining them confirms both the presence and functional activity of targets, providing mechanistic evidence that strengthens regulatory submissions for biologics and ATMPs.
How can AI improve molecular assay development?
AI aids assay selection and performance optimization by automating variant caller tuning, multiplexing design, and quality control flagging, particularly for complex regulatory alignment challenges.



